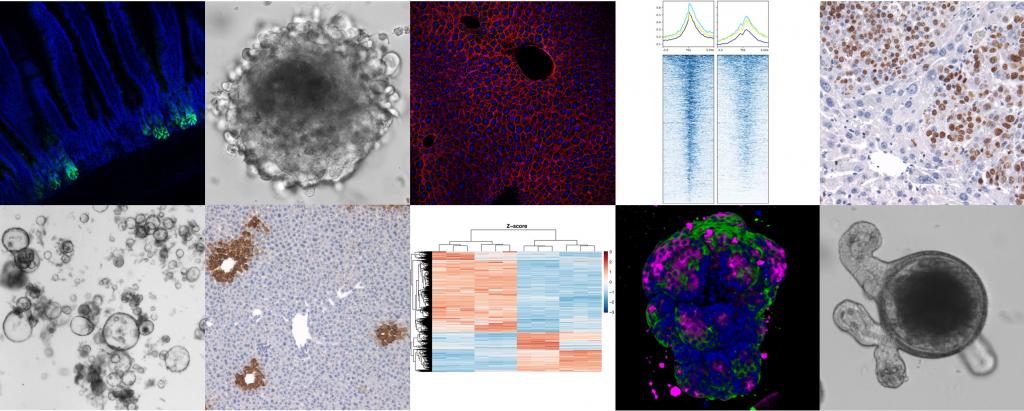
Overview
Adult tissue homeostasis is maintained by the accurate integration of the different signaling pathways required to preserve cell identity. Stem cell maintenance and cell lineage determination are regulated by the coordinated activity of specific signaling pathways that converge on chromatin to activate and repress well-defined transcriptional programs. By shaping the epigenomic landscape of cells, transcription factors and chromatin modifying complexes orchestrate these processes. During tumor formation driver oncogenic mutations lead to the emergence of a progenitor-like state characterized by a change in cell identity and an extensive transcriptional reprograming. This reprograming is the result of widespread rearrangement in the repertoire of active functional elements, due to alterations at different levels in their regulation: mutation in their sequence, in the expression of different transcription factors, or aberrant activity of chromatin modifying enzymes, often mutated in cancer.
In our laboratory we investigate how external and internal stimuli are integrated at the chromatin level. In particular, by using a comprehensive approach, we aim to describe the role of chromatin modifying enzymes and transcription factors in the maintenance of adult stem cells and in doing so characterize the transcriptional mechanisms and the chromatin associated events involved in tumor formation and metastatic progression. Through the extensive characterization of these mechanisms our ultimate goal is the identification of putative targets for novel pharmacological approaches.
Research Projects
- Characterize the transcriptional and epigenetic events driving liver regeneration and liver tumors formation and progression.
- Characterize the epigenetic and mutagenic events required for dissemination and growth of lung metastasis from hepatocellular carcinoma.
- Exploit patients-derived tumor organoids as models for pre-clinical studies.
Group members
- Fulvio Chiacchiera, PI
- Erna Marija Meškytė, Post-doc
- Davide Bressan, PhD student
- Elisa Ferracci, PhD student
- Doina Sirbu, undergraduate student
- Emma Zocca, undergraduate student
- Chiara Magro, undergraduate student
PhD student and undergraduate student positions are available. Please contact fulvio.chiacchiera [at] unitn.it (Fulvio Chiacchiera) directly
Collaborations
- Prof. Diego Pasini, European Institute of Oncology (IEO), Italy
- Dr Giovanni Sorrentino, University of Trieste, Italy
- Prof. Mattia Barbareschi S.Chiara Hospital and University of Trento, Italy
- Dr Roberto Gherzi, IRCCS Ospedale Policlinico San Martino, Genova
- Dr. Piergiorgio Amendola, Dompè Farmaceutici s.p.a, Italy
Funding
Bando: PRIN 2022 (D.D. 104/22)
Revealing The Contribution Of Nuclear Mechanics In Non-Alcoholic Fatty Liver Disease Progression
Fulvio Chiacchiera, Responsabile di Unità
Codice Protocollo: 2022PWKZXE CUP: E53D23013290006
Progetto: PRIN/PNRR
Defining the Mechanisms that Connect Oncogenic BAP1 Loss of Function Mutations to Environmental Stresses in Cholangiocarcinoma
Fulvio Chiacchiera, Responsabile di Unità
Codice progetto: P2022YP3HZ CUP: E53D23015300001

- Worldwide Cancer Research
- Italian Association for Cancer Research (AIRC)
Awards
2016, Begnudelli Award, Pezcoller-AACR Foundation, Trento, Italy
Selected publications
D'Ambrosio A*, Bressan D*, Ferracci E*, Carbone F, Mulè P, Rossi F, Barbieri C, Sorrenti E, Fiaccadori G, Detone T, Vezzoli E, Bianchi S, Sartori C, Corso S, Fukuda A, Bertalot G, Falqui A, Barbareschi M, Romanel A, Pasini D, Chiacchiera F. (2024) Increased genomic instability and reshaping of tissue microenvironment underlie oncogenic properties of Arid1a mutations. Science Advances 15;10(11):eadh4435
Briata P, Mastracci L, Zapparoli E, Caputo L, Ferracci E, Silvestri A, Garuti A, Hamadou MH, Inga A, Marcaccini E, Grillo F, Bucci G, Puri PL, Beznoussenko G, Mironov A, Chiacchiera F*, Gherzi R.* (2023) LncRNA EPR regulates intestinal mucus production and protects against inflammation and tumorigenesis. Nucleic Acids Res. doi: 10.1093/nar/gkad257. Online ahead of print.
Biferali B, Bianconi V, Perez DF, Kronawitter SP, Marullo F, Maggio R, Santini T, Polverino F, Biagioni S, Summa V, Toniatti C, Pasini D, Stricker S, Di Fabio R, Chiacchiera F, Peruzzi G, Mozzetta C. (2021) Prdm16-mediated H3K9 methylation controls fibro-adipogenic progenitors identity during skeletal muscle repair. Science Advances 7(23):eabd9371.
Bisso A, Filipuzzi M, Gamarra Figueroa GP, Brumana G, Biagioni F, Doni M, Ceccotti G, Tanaskovic N, Morelli MJ, Pendino V, Chiacchiera F, Pasini D, Olivero D, Campaner S, Sabò A, Amati B. (2020) Cooperation Between MYC and β-Catenin in Liver Tumorigenesis Requires Yap/Taz. Hepatology 72(4):1430-1443.
Pivetti S., Fernandez-Perez D., D’Ambrosio A., Barbieri C.M., Manganaro D., Rossi A., Barnabei L., Zanotti M., Scelfo A., Chiacchiera F.* and Pasini D.* (2019). Loss of PRC1 activity in different stem cell compartments activates a common transcriptional program with cell-type dependent outcomes. Science Advances 5(5): eaav1594.
Chiacchiera F., Pasini D., (2017) Control of Adult Intestinal Identity by the Polycomb Repressive Machinery. Cell Cycle 16(3):243-244.
Chiacchiera F.*, Rossi A., ShriGanesh J., Zanotti M., Pasini D.* (2016) PRC2 preserves intestinal progenitors and restricts secretory lineage commitment. Embo j. 35(21):2301-2314.
Rossi A., Ferrari K.J., Piunti A., ShriGanesh J., Chiacchiera F., Mazzarella L., Scelfo A., Pelicci P.G., Pasini D. (2016) Maintenance of leukemic cell identity by the activity of the polycomb complex PRC1 in mice. Science Advances 2(10): e1600972.
Chiacchiera F., Rossi A., ShriGanesh J., Piunti A., Scelfo A., Ordonez- Morànx P., Huelsken J., Koseki H., Pasini D. (2016) Polycomb Complex PRC1 preserves intestinal stem cell identity by sustaining Wnt/β-Catenin transcriptional activity. Cell Stem Cell, 18(1):91-103.