Overview
Vaccination is one of the most effective intervention in the history of medicine. Devastating diseases caused by various infectious agents have been completely or almost completely eliminated thanks to the introduction of vaccination practice. However, there are a number of infectious diseases for which vaccines do not exist yet.
Furthermore, chronic diseases such as cancer are still awaiting valid solutions to turn vaccination into an effective prophylactic and/or therapeutic intervention. 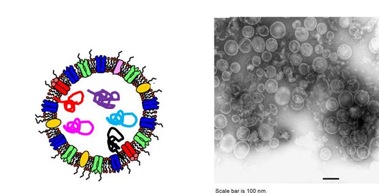
The development of vaccines against recalcitrant infections and cancer will depend on the identification of highly specific antigens to be formulated with potent adjuvants capable of eliciting effective immune responses. The main objective of our Laboratory is to create novel vaccine platforms based on the exploitation of the Outer Membrane Vesicles (OMVs) naturally released by Gram-negative bacteria. Because of the intrinsic capacity of OMVs to potently stimulate innate immunity and thanks to the possibility of OMVs to be manipulated with heterologous antigens the vesicles have the potential to become the scaffold of highly innovative and effective vaccines.
Research directions
The SSV Laboratory was set up in 2014 when the PI was assigned an Advanced ERC Grant from the European Community. The main activities of the SSV Laboratory can be summarized as follows:
-
Genome editing of Escherichia coli to create a new strain specialized in OMVs production
OMVs carry a conspicuous number of endogenous proteins mainly belonging to outer membrane and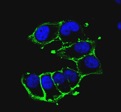 periplasmic compartments. It is expected that the alteration of the protein repertoire of OMVs could profoundly affect the immune response elicited in mammals both in qualitative and quantitative terms. Innovative approaches for genome editing are being applied to selectively remove/add genes and gene pathways to ultimately create an E. coli strain capable of releasing abundant quantities of OMVs carrying a minimal amount of endogenous proteins and novel immune-stimulatory components.
periplasmic compartments. It is expected that the alteration of the protein repertoire of OMVs could profoundly affect the immune response elicited in mammals both in qualitative and quantitative terms. Innovative approaches for genome editing are being applied to selectively remove/add genes and gene pathways to ultimately create an E. coli strain capable of releasing abundant quantities of OMVs carrying a minimal amount of endogenous proteins and novel immune-stimulatory components. -
Development of novel genetic tools to decorate OMVs with foreign antigens
The main objective of this activity is to find gene technology strategies to deliver the desired protein antigens to the OMV compartments. Such technologies will include gene fusions, domain engineering and grafting of transport machineries. -
Development of novel OMV-based vaccines against bacterial pathogens and cancer
OMVs are being decorated with selected bacterial and cancer antigens and immunogenicity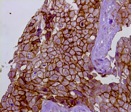 and protective activity of OMVs are being evaluated in vitro and in vivo. In particular, the mechanisms of stimulation of innate immunity is being analyzed using cell lines engineered with specific Patterns Recognition Receptors (PRRs). Furthermore, the elicitation of antibodies, CD4 and CD8 T cells will be evaluated in appropriate animal models. Finally, mouse models for infectious diseases and cancer will be set-up and used to establish the protective properties of OMV-based vaccines and to correlate protection to humoral and cell-mediated immunity.
and protective activity of OMVs are being evaluated in vitro and in vivo. In particular, the mechanisms of stimulation of innate immunity is being analyzed using cell lines engineered with specific Patterns Recognition Receptors (PRRs). Furthermore, the elicitation of antibodies, CD4 and CD8 T cells will be evaluated in appropriate animal models. Finally, mouse models for infectious diseases and cancer will be set-up and used to establish the protective properties of OMV-based vaccines and to correlate protection to humoral and cell-mediated immunity.
Group members
- Guido Grandi, PI
- Laura Fantappiè, postdoctoral fellow
- Francesca Zerbini, postdoctoral fellow
- Carmela Irene, postdoctoral fellow
- Luisa Ganfini, PhD student
- Michele Tomasi, PhD student
Collaborations
- Renata Grifantini, Alberto Grandi, Matteo Parri, Piero Pileri, Susanna Campagnoli, Externautics SpA, Milan, Italy
- Olaf Schneewind, University of Chicago, Chicago, Illinois, USA
- Olivera (Olja) J. Finn, University of Pittsburgh School of Medicine, Pittsburgh, PA, USA
- Rajiv Khanna, QIMR Berghofer, Brisbane, Australia
Selected publications
F. Bagnoli, M.R. Fontana, E. Soldaini, R.P. Mishra, L. Fiaschi, E. Cartocci, V. Nardi-Dei, P. Ruggiero, S. Nosari, M.G. De Falco, G. Lofano, S. Marchi, B. Galletti, P. Mariotti, M. Bacconi, A. Torre, S. Maccari, M. Scarselli, C.D. Rinaudo, N. Inoshima, S. Savino, E. Mori, S. Rossi-Paccani, B. Baudner, M. Pallaoro, E. Swennen, R. Petracca, C. Brettoni, S. Liberatori, N. Norais, E. Monaci, J. Bubeck Wardenburg, O. Schneewind, D.T. O'Hagan, N.M. Valiante, G. Bensi, S. Bertholet, E. De Gregorio, R. Rappuoli and G. Grandi* (2015). Vaccine composition formulated with a novel TLR7-dependent adjuvant induces high and broad protection against Staphylococcus aureus. Proc Natl Acad Sci U S A. 112:3680-5.
E. Altindis, R. Cozzi, B. Di Palo, F. Necchi, R.P. Mishra, M.R. Fontana, M. Soriani, F. Bagnoli, D. Maione, G. Grandi* and S. Liberatori (2015). Protectome analysis: a new selective bioinformatics tool for bacterial vaccine candidate discovery. Mol Cell Proteomics. 14:418-29.
L. Fantappiè, M. de Santis, E. Chiarot, F. Carboni, G. Bensi, O. Jousson, I. Margarit and G. Grandi* (2014). Antibody-mediated immunity induced by engineered Escherichia coli OMVs carrying heterologous antigens in their lumen. J Extracell Vesicles. 3. doi: 10.3402/jev.v3.24015.
E. Bartolini, E. Ianni, E. Frigimelica, R. Petracca, G. Galli, F. Berlanda Scorza, N. Norais, D. Laera, F. Giusti, A. Pierleoni, M. Donati, R. Cevenini, O. Finco, G. Grandi* and R. Grifantini (2013). Recombinant outer membrane vesicles carrying Chlamydia muridarum HtrA induce antibodies that neutralize chlamydial infection in vitro. J Extracell Vesicles. 2: 20181.
P.R. Dormitzer, G. Grandi and R. Rappuoli (2012) Structural vaccinology starts to deliver. Nat Rev Microbiol. 10:807-813.
O. Finco, E. Frigimelica, F. Buricchi, R. Petracca, G. Galli, E. Faenzi, E. Meoni, A. Bonci, M. Agnusdei, F. Nardelli, E. Bartolini, M. Scarselli, E. Caproni, D. Laera, L. Zedda, D. Skibinski, S. Giovinazzi, R. Bastone, E. Ianni, R. Cevenini, G. Grandi* and R. Grifantini (2011). Approach to discover T and B cell antigens of intracellular pathogens applied to the design of Chlamydia trachomatis vaccines. Proc. Natl. Acad. Sci. U S A. 108:9969-9974.
A. Nuccitelli, R. Cozzi, L.J. Gourlay, D. Donnarumma, F. Necchi, N. Norais, J.L. Telford. R. Rappuoli, M. Bolognesi, D. Maione, G. Grandi* and C.D. Rinaudo (2011). A structure-based approach to rationally design a chimeric protein for an effective vaccine against Group B Streptococcus infections. Proc. Natl. Acad. Sci. U S A. 108:10278-10283.
